Background
Rhesus D genotyping with cell-free fetal DNA currently is used throughout the world. Although this technique has spread rapidly, its optimal use is still a matter of debate. This screening test has been introduced mainly for the treatment of RhD-negative pregnant women during the third trimester of pregnancy, thereby avoiding systematic anti-D prophylaxis, yet such a strategy has proved cost-ineffective. Publications reporting on fetal RHD genotyping with cell-free DNA in maternal plasma, specifically during the first trimester of pregnancy, are scarce in the scientific literature.
Objective
This study sought to assess the performance of noninvasive fetal Rhesus D genotyping in the first trimester of pregnancy with a single-exon real-time polymerase chain reaction assay.
Study Design
This was a retrospective observational multicenter study. Cell-free fetal DNA was extracted from maternal blood of both nonimmunized and immunized women at 10–14 weeks of gestation. RHD sequence was determined by quantitative polymerase chain reaction, with amplification of exon 10. Results were compared with RhD phenotype data that were obtained by cord blood sampling of neonates.
Results
In total, 416 serum samples from RhD-negative pregnant women were collected during the first trimester of pregnancy. The test’s overall sensitivity and specificity were 100% (95% confidence interval, 96.9–100.0) and 95.2% (95% confidence interval, 90.5–97.6), respectively. The negative and positive predictive values were 99.8% (95% confidence interval, 94.9–100.0) and 97.1% (95% confidence interval, 94.2–98.6), respectively. Fetal RHD status was inconclusive in 9 cases (2.2%).
Conclusion
Noninvasive fetal RHD determination by single-exon quantitative polymerase chain reaction during the first trimester of pregnancy exhibits high accuracy.
Since 1997 when Lo et al reported detecting cell-free fetal DNA in the plasma of pregnant women, numerous applications have been developed, which has led to an entirely new era for noninvasive prenatal diagnosis. As a major application, noninvasive rhesus D (RhD) genotyping has revolutionized the determination of fetal RHD status for RhD-negative women. Before RHD genotyping by polymerase chain reaction (PCR), fetal RHD status was ascertained after an invasive procedure (ie, chorionic villus sampling or amniocentesis). At the present time, the assay is based on the detection of RHD gene sequences in the plasma of RhD-negative women. This assay was proposed initially for small cohorts of RhD-immunized women to exclude any risk of RhD incompatibility or, conversely, adapt patient care for pregnancies at risk of hemolytic disease and fetal anemia. Thereafter, RHD genotyping has been assessed in large populations of nonimmunized women, with promising results.
In 2002, the first noninvasive fetal RHD genotyping test was introduced by our team in routine practice in France. Solely based on the detection of RHD gene exon 10 by real-time PCR, this assay, which initially was assessed in a cohort of 102 first-trimester pregnant women, exhibited perfect accuracy (100% sensitivity and specificity). Next, the assay was evaluated in a cohort of 283 women for whom reliable fetal RHD genotype determination could again be achieved with 100% accuracy.
Here we have reported the results that pertain to our extended experience in the noninvasive prenatal determination of fetal RHD status that is proposed routinely in the first pregnancy trimester.
Material and Methods
Study population
In total, 416 RhD-negative pregnant women were enrolled retrospectively in 5 obstetrics departments in the Paris area (American Hospital of Paris, Institut Mutualiste Montsouris, Centre Hospitalo-Universitaire Necker-Enfants Malades, Centre Hospitalo-Universitaire Antoine Béclère, and Institut Hospitalier Franco-Britannique) between January 2007 and December 2012. All blood samples were obtained at 10–14 weeks of gestation as part of a routine consultation or specific prenatal diagnosis consultation in relation to previous RhD immunization. Medical indications for noninvasive fetal RHD determination in RhD-negative women were (1) systematic screening of nonimmunized RhD-negative women, to guide anti-D administration after an invasive procedure (chorionic villus sampling or amniocentesis) if needed or (2) events at risk for RhD immunization (abdominal trauma, miscarriage, or metrorrhagia).
Samples
Blood samples (7 mL) were collected into Vacutainer SST tubes (Becton Dickinson, Meylan, France). Immediately after clotting, serum was obtained by centrifugation at 3000 g for 10 minutes at 4°C, aliquoted, and stored at –80°C. If not treated immediately, the blood sample was kept at room temperature for a maximum of 72 hours before being processed. Anti-D immunoglobulins were not administered to patients whose fetus was found to be RhD-negative.
Real-time PCR for RHD gene
Detection of sequences derived from the human RHD gene was performed, as described elsewhere ; the mouse GALT gene was used as a positive control of DNA extraction. Briefly, as a tracer for DNA extraction and amplification steps, a low amount (250 pg) of mouse DNA (Sigma, Grenoble, France) was added to each patient’s sample (1 mL of serum) immediately before DNA extraction. Total DNA was then extracted with the use of the PCR Template Preparation Kit (Roche Diagnostics, Meylan, France), and the adsorbed DNA was eluted with 50 μL of elution buffer, 10 μL of which was used per PCR reaction. Amplification was carried out in a Light-Cycler V2.0 instrument (Roche Diagnostics). PCR reactions were performed with the use of the FastDNAMaster Hybridization Probes Kit (Roche Diagnostics) in a final volume of 20 μL, with 0.5 mmol/L of each primer ( Table ), 0.25 mmol/L of each probe (Sigma), 1.25 units of uracil-DNA glycosylase (Biolabs, Saint-Quentin en Yvelines, France), and 4.75 mmol/L of magnesium chloride. After an initial 1-minute incubation at 50°C, a first denaturation step of 8 minutes at 95°C was followed by amplification performed for 50 cycles of denaturation (95°C, 10 seconds, ramping rate 20°C/sec), annealing (56°C, 10 seconds, ramping rate 20°C/sec), and extension (72°C, 20 seconds, ramping rate 2°C/sec). Each sample was treated twice for DNA extraction, and the RHD assay was performed in duplicate on each DNA extract. During each run, sera that were obtained from women carrying an RhD-positive or RhD-negative fetus were used as positive and negative controls. RHD reaction was considered to be positive when a fluorescent signal was detected for the RHD gene and to be negative when a signal was detected for the mouse GALT gene only. The results were considered inconclusive when the crossing point value (fluorescent signal) that was observed for the RHD reaction appeared much earlier than for the positive control. In that specific case, it means that the amount of RHD sequences that are detected in maternal serum is too large to be of fetal origin. Based on these findings, it can be concluded that RHD gene sequences are present in the maternal genome, which suggests the presence of a mother variant of RHD gene. Definitive RhD-negative genotype was considered established when all RHD PCR reactions were negative, and RHD -positive genotype was considered established when at least 3 of 4 PCR reactions were positive.
| Variable | Target | Sequence (5′-3′) | Modification |
|---|---|---|---|
| Primer name | |||
| RHE10A | Human RHD gene | GCCTGCATTTGTACGTGAGA | None |
| RHE10B | Human RHD gene | CAAAGAGTGGCAGAGAAAGGA | None |
| IS11 | Mouse GALT gene | GCGCTTCCCGAGGTACACTAT | None |
| IS12 | Mouse GALT gene | ATGTCACATCTGCCCGAACTCC | None |
| Probe name | |||
| RHE10C | Human RHD gene | GCAGGCACTGGAGTCAGAGAAAA | 5′LCRed640 3′Ph |
| RHE10D | Human RHD gene | TGACAGCAAAGTCTCCAATGTTCG | 3′FITC |
| IS9 | Mouse GALT gene | TGGTGATCCTGCCGTTTCCTTGTCTT | 5′LCRed705 3′Ph |
| IS10 | Mouse GALT gene | GCCCTGATGTGGTCACAGTCAAGCA | 3′FITC |
The results were compared with those obtained by RhD serology of the neonate cord blood sample.
Data and statistical analyses
For each woman who was included in the study, the following data were collected: gestational age at noninvasive RHD determination, indication for RHD determination, results of real-time PCR for RHD gene, neonatal RhD determination (cord blood sampling) as gold standard, antenatal and postnatal screenings for irregular antibodies, antenatal or postnatal anti-D administration, and geographic origin of both parents.
Data were analyzed with JMP 10 Statistical Discovery software (SAS Institute, Cary, NC). All estimates were presented with 95% confidence intervals (CI).
Details of ethics approval
This observational retrospective study was approved by the institutional review board of the French college of obstetricians and gynecologists (CEROG OBS 2013-02-04 R1). All data were deidentified to ensure patient privacy and confidentiality. According to the French regulation regarding prenatal diagnosis, written informed consent was obtained from all patients.
Results
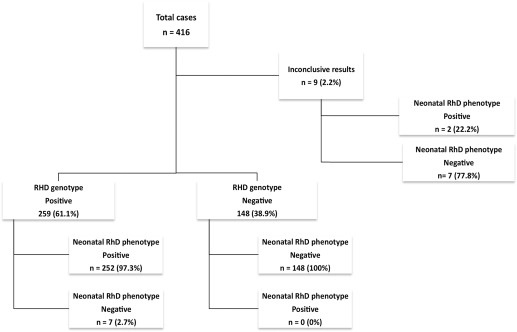
Among the 416 RhD-negative women who were tested for fetal RHD status, the mean gestational age at blood sampling was 13.1 ± 1.0 weeks. Overall 25 procedures (6.0%) were performed at 10–11 weeks of gestation. Most women (90.6%) were tested as part of systematic screening; 22 women (5.3%) were tested before an invasive procedure, and 17 women (4.1%) were tested because of existing RhD immunization. Among women who were tested for fetal RHD status, 48 women (11.5%) were of African or Caribbean origin. Neonatal RhD status was negative in 162 cases (38.9%) and positive in 254 cases (61.1%).
In this study cohort, fetal RHD status was inconclusive in 9 cases (2.2%; Figure ), with 6 of them being of African or Caribbean origin. Concerning these inconclusive results, 2 newborn infants were determined to be RhD-positive, and 7 infants were determined to be RhD-negative (based on cord blood sera).